Phosphate metabolism and signaling in plants
Phosphorus is essential for life as we know it. It forms the foundation of DNA, lipids, and proteins, and serves as the energy currency (in the form of ATP) in all living cells. Organisms have developed advanced mechanisms to sense, acquire, transform, and store phosphorus, primarily as inorganic phosphate. For plants, which absorb it directly from the soil, phosphate is a growth-limiting nutrient that is often provided in large quantities as fertilizer to meet the high-yield needs of modern agriculture. Although we have made strides in understanding how plants respond to phosphate starvation, the precise mechanism by which phosphate levels are “measured” remains unclear. Our lab focuses on advancing the understanding of phosphate sensing, signaling, and storage by studying various phosphate-containing molecules, including inorganic polyphosphates and inositol pyrophosphates (Lorenzo-Orts et al., New Phytol, 2020).
Inorganic polyphosphate – an ubiquitous but enigmatic phosphate polymer
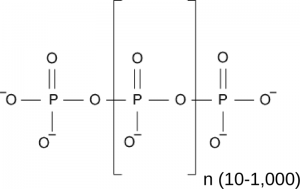
One rather poorly understood storage form of inorganic phosphate in all pro- and eukaryotic cells is inorganic polyphosphate, a linear chain of orthophosphate units linked by phosphoanhydride bonds (Figure 1). In bacteria, polyP is generated from ATP by polyP kinase (Sylvie Kornberg, BBA, 1957) and serves important physiological roles (Kornberg et al., Ann Rev Biochem, 1999) . Inorganic polyphosphate kinases however are absent from eukaryotic genomes, and thus it was unknown for a long time how eukaryotes would synthesize inorganic polyphosphates.
A vacuolar integral membrane protein complex synthesizes polyP in yeast
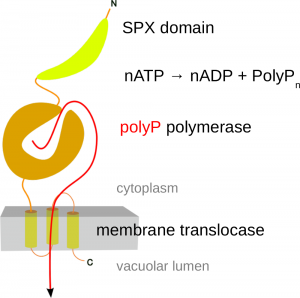
Through serendipity, we discovered the first eukaryotic enzyme capable of synthesizing inorganic polyphosphate. (Hothorn et al., Science, 2009). It was found to be an integral membrane protein complex located in the vacuolar membrane, rather than a soluble enzyme. This complex, known as the Vacuolar Transporter Chaperone (VTC) Complex (a name chosen because its exact function is still unclear), is composed of at least three distinct subunits. Each subunit contains a transmembrane domain (as detailed below), and two of the subunits (Vtc2/3 and Vtc4) also have soluble portions that face the yeast cytoplasm (Mueller et al., EMBO J, 2002) (Figure 2). Through X-ray protein crystallography, we were able to show that the soluble regions of VTC contain a tunnel-shaped triphosphate tunnel metalloenzyme (TTM) domain. Our findings revealed that the Vtc4 TTM domain could bind and hydrolyze ATP in the presence of a manganese metal cofactor. After dialyzing the tunnel domain in a solution with ATP, we obtained crystals of Vtc4. To our surprise, these crystals did not contain ATP bound in the catalytic cleft, but instead, a long phosphate polymer. (Figure 3) (Hothorn et al., Science, 2009)
.
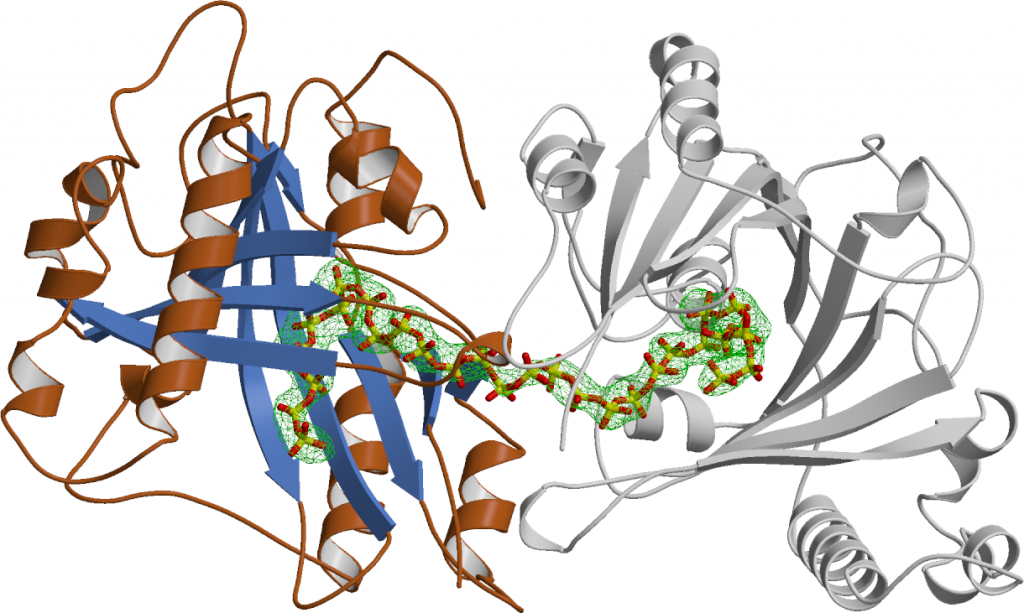
VTC turned to be the long sought-for polyphosphate polymerase in yeast. Yeast cells contain high concentrations of polyphosphate in their vacuole. Point-mutations in the VTC transmembrane domain inhibit translocation of the polymer into this storage compartment. Upon inactivation of polyphosphate polymerisation in VTC, yeast mutants cannot survive under phosphate limiting conditions and have multiple cellular defects. A single particle cryo electron microscopy structure of the entire VTC complex has now been resolved (Guan et al., Nat Commun, 2023). There is mounting evidence that inorganic polyphosphates are important storage forms of phosphate in eukaryotic cells. They may also play roles in ion homeostasis and energy metabolism.
The metabolism and cellular functions of polyphosphates in higher eukaryotes
The presence of inorganic polyphosphates has been well documented in bacteria, unicellular eukaryotes and animals. We are, however, still not certain of its existence in plants (Lorenzo-Orts et al., New Phytol, 2020). Unfortunately, the VTC complex is only conserved in unicellular eukaryotes and not present in plants (or in animals, for that matter). We thus aimed at identifying other polyphosphate metabolizing enzymes in the model plant Arabidopsis thaliana. We also tried to localize inorganic polyphosphates in plants and to understand their cellular and physiological functions. This work was supported by an ERC starting grant.
Arabidopsis contains AtTTM3, a soluble enzyme with structural homology to yeast Vtc4
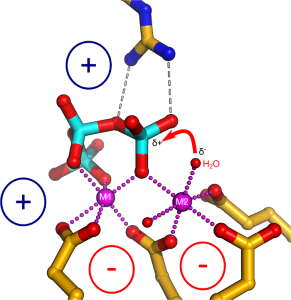
We employed a structure-guided approach to identify Arabidopsis proteins resembling Vtc4. The Arabidopsis genome contains three TTM proteins (AtTTM1-3), with TTM3 being a small soluble enzyme that shares a high structural similarity with Vtc4 (Martinez, Truffault & Hothorn, JBC, 2015). However, postdoctoral fellow Jacobo Martinez-Font discovered that, rather than polymerizing inorganic polyphosphate from ATP, AtTTM3 efficiently hydrolyzes short-chain polyphosphates in vitro. This raised the question: why does the catalytic activity differ between the yeast and plant enzymes? Jacobo investigated the catalytic mechanisms of TTM enzymes from plants, yeast, bacteria, and humans, analyzing both their mechanisms and structures in detail. He found that most TTM enzymes utilize a two-metal catalytic mechanism to hydrolyze their substrates (Figure 4).
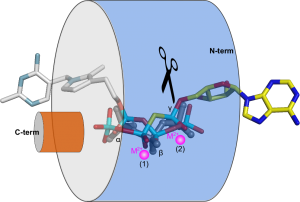
Vtc4 however, uses only one catalytic metal, with the second metal binding site modified in such a way, that it can harbor a phosphate polymer to which the ATP γ-phosphate can be transferred to (Martinez, Truffault & Hothorn, JBC, 2015). Jacobo’s work also provided insights into how TTM enzymes are capable of performing such a diverse range of enzymatic reactions (including adenylate cyclase, thiamine triphosphatase, tripolyphosphatase, and others). He discovered that different TTM enzymes allow their substrates to enter the tunnel domain from opposite sides. This structural flexibility enables the core catalytic center to remain in a consistent position while the enzymes can generate either phosphate or pyrophosphate leaving groups (Figure 5). Additionally, Jacobo resolved the structures of AtTTM1/2 and demonstrated that these enzymes function as phosphotransferases, transferring the γ-phosphate of an NTP substrate to an unknown acceptor molecule that may be bound to the catalytically inactive TTM domain (Pesquera, Martinez et al., JBC, 2022).
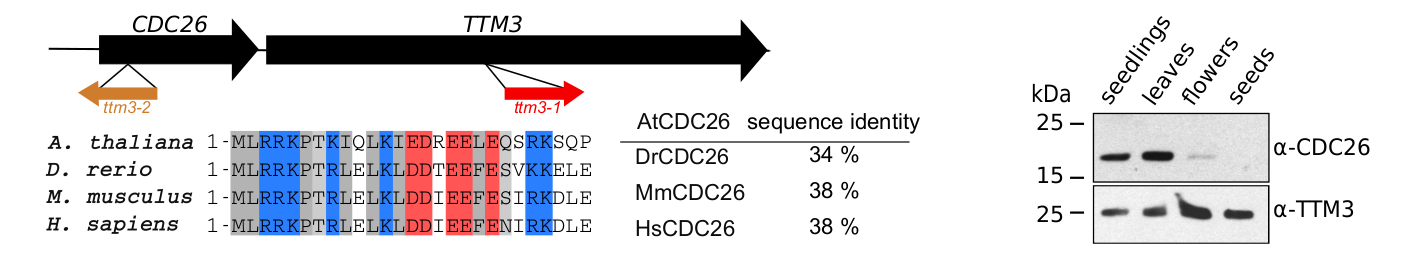
Our characterization of AtTTM3 as a specific short-chain polyphosphatase conserved in the entire green lineage prompted us to investigate the physiological functions of the enzyme in Arabidopsis. Postdoctoral fellow Janika Witthoeft and PhD student Laura Lorenzo-Orts investigated ttm3 loss-of-function alleles that displayed strong phenotypes during embryo development. Laura then found that AtTTM3 is transcribed into a bicistronic mRNA that encodes for both AtTTM3 and the cell division protein 26 (CDC26) (Lorenzo-Orts et al., Nature Plants, 2019) (Figure 6). Not only did Laura identify an extremely rare eukaryotic bicistronic mRNA, she also discovered CDC26, which was thought to be absent in the green lineage. Laura established that CDC26 is in fact a subunit of the anaphase promoting complex/cyclosome (APC/C), and that AtTTM3 coordinates AtCDC26 translation by recruiting the transcript into polysomes (Lorenzo-Orts et al., Nature Plants, 2019). While there seems to be no biochemical connection between the two proteins, this bicistronic configuration is conserved across the plant lineage suggesting an important regulatory function.
CHAD domains are inorganic polyphosphate binding modules
Laura next identified other inorganic polyphosphate-related proteins in plants. Intriguingly, she found that in at least one plant, Ricinus communis, a CHAD-domain containing protein that shares high structural and sequence similarly to the CHAD domains present in bacteria and archae (Lorenzo‐Orts et al., Life Sci Alliance, 2019) (Figure 7).
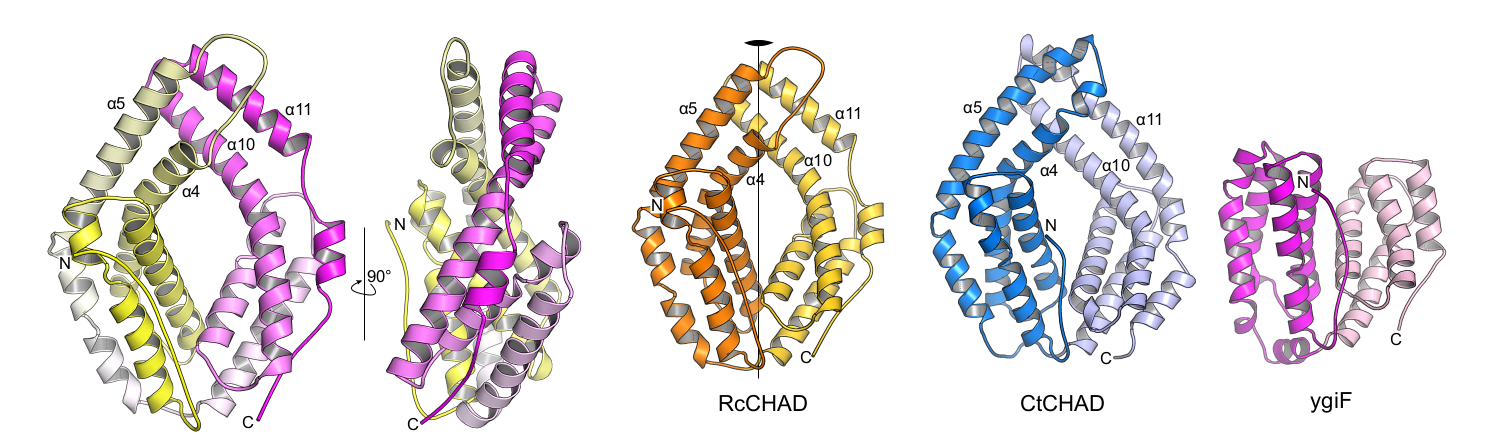
Next Laura showed that CHAD domains specifically bind inorganic polyphosphates with high affinity and selectivity and solved a structure of a CHAD domain – polyP complex (Figure 8).
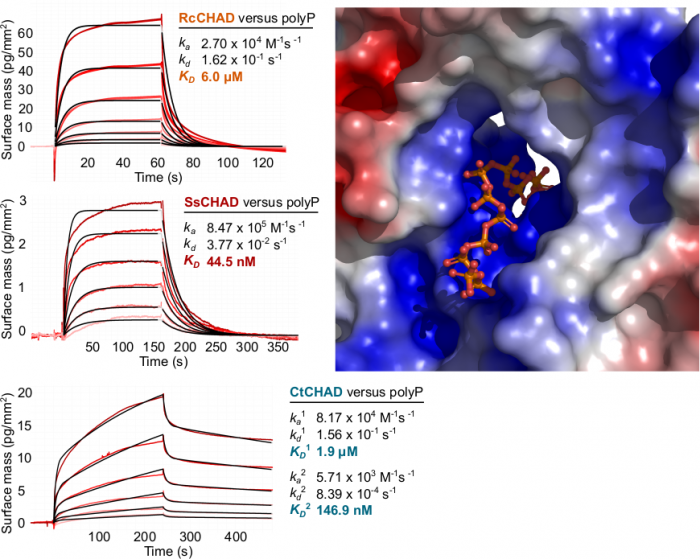
Taken together CHAD domains of previously unknown function are conserved inorganic polyphosphate binding modules present in the different kingdoms of life. Its singularity in the plant kingdom suggests that RcCHAD might have been acquired by Ricinus via horizontal gene transfer from a soil‐living bacterium. Whether it acquired a novel function or still binds inorganic polyphosphates in Ricinus remains to be investigated. We could exploit the inorganic polyphosphate binding specificity of CHAD domains to use them as probes for the detection of inorganic polyphosphates in plant cells. Notably, fluorescent protein fused CHAD domains localize to the nucleolus, suggesting that a inorganic polyphosphate store may exist in this subcellular compartment (Lorenzo‐Orts et al., Life Sci Alliance, 2019).
Detection methods for inorganic polyphosphates in plant cells and tissues
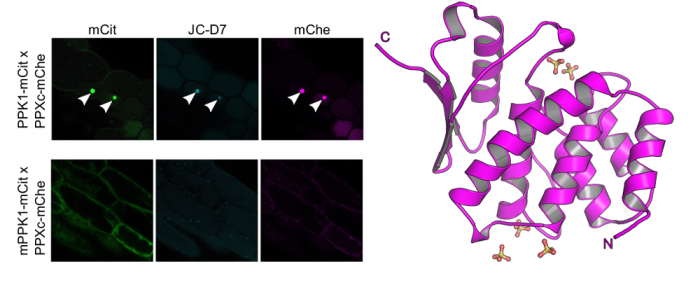
We complemented our search for bona fide inorganic polyphosphate binding- or metabolizing proteins in Arabidopsis by looking for stores of this enigmatic polymer in living plant cells and tissues. To this end postdoctoral fellow Jinsheng Zhu used a fluorescent dye (JC-D7), which specifically binds inorganic polyphosphate but not other phosphate-containing molecules, such as nucleic acids, nucleotides or phosphate salts (Angelova et al., ACS Chem Biol, 2014). While he was able to visualize inorganic polyphosphate granules in Arabidopsis cells expressing a bacterial inorganic polyphosphate kinase, he could not detect inorganic polyphosphates in wild‐type Arabidopsis plants, nor in other monocotyledonous or dicotyledonous plants (Figure 9) (Zhu et al., Plant J, 2019).
Taken together, we located inorganic polyphosphate binding proteins and metabolic enzymes in plants. However, we failed to detect significant inorganic polyphosphate stores in higher plants and could not define clear loss-of-function phenotypes for the inorganic polyphosphate phosphatase TTM3. This suggests that inorganic polyphosphates in plants either don’t exist or do not accumulate to high levels. They may only be present in specific cell types or organs, or may be produced in response to certain environmental stimuli not replicated in our in our plant growth chambers.
SPX domains are inositol pyrophosphate sensor domains controlling phosphate homeostasis in eukaryotic cells
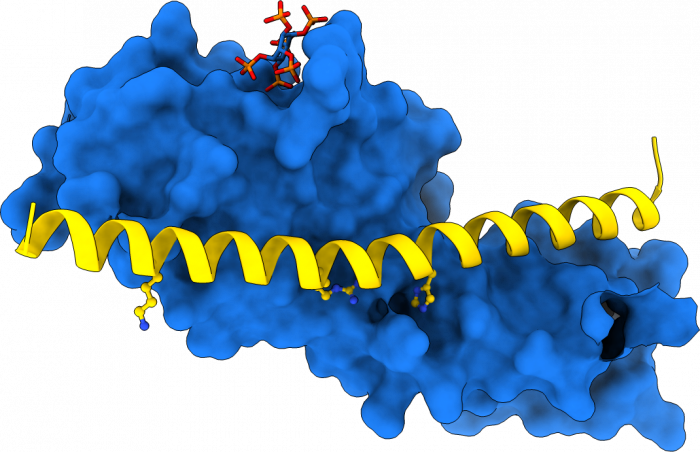
In parallel to our analysis of plant inorganic polyphosphate metabolism, PhD student Rebekka Wild investigated the molecular functions of the N-terminal SPX domains in the VTC complex and in other eukaryotic proteins (Wild et al., Science, 2016). SPX domains are small, soluble domains found in fungi, plants and animals. They can exist as stand-alone modules but are often located at the N-termini of proteins involved in phosphate uptake, transport, storage, metabolism or signaling. We determined structures of fungal and human SPX domains from crystals obtained by carrier driven crystallization (Wild & Hothorn, Protein Science, 2016). The different structures revealed a new fold with two long a-helices, connected by linkers of variable size. These core helices and two smaller C-terminal helices together form a 3-helix bundle, which is preceded by an flexible N-terminal helical hairpin (Figure 10). Many invariant lysine residues, which represent sequence fingerprints for SPX domains, are clustered in proximity to the N-terminal hairpin structure. A combination of genetic and biochemical experiments in yeast and Arabidopsis by the groups of Andreas Mayer and Yves Poirier, University of Lausanne, revealed that this basic surface represents a docking platform for inositol pyrohosphosphates (PP-InsPs), signaling molecules with poorly characterized cellular functions (Figure 10).
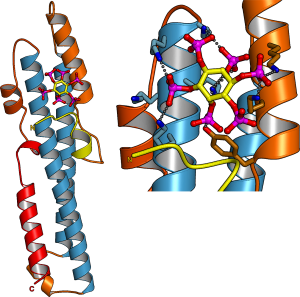
The concentrations of PP-InsP are known to change in cells, depending on whether they are supplied with sufficient amounts of inorganic phosphate or whether they experience phosphate starvation. Rebekka could demonstrate that SPX domains, when bound to PP-InsPs can interact with other proteins. In plants, one such interaction partner is a transcription factor, which is switched on under phosphate starvation to induce expression of genes that allow the plant to respond to the lack of this important nutrient. Under normal growth conditions, when PP-InsP levels are high, the transcription factor is kept inactive by forming a PP-InsP-dependent complex with a plant SPX domain. Under phosphate starvation, the lack of PP-InsP leads to dissociation of the transcription factor – SPX domain complex, thereby enabling the transcription factor to transcribe its target genes. Taken together, Rebekkas work defined SPX domains as cellular receptor for PP-InsPs, which control phosphate uptake, transport, storage, metabolism and signaling in fungi, plants and animals.
Two bifunctional inositol pyrophosphate kinases/phosphatases regulate plant phosphate homeostasis
The next logical question was how inositol pyrophosphate signaling molecules are generated in plants and why their concentrations fluctuate in response to phosphate availability. Postdoctoral fellows Jinsheng Zhu and Kelvin Lau addressed this question by employing a combination of plant genetics, cell biology, and in vitro enzymology. Jin characterized two homologous inositol pyrophosphate kinases/phosphatases, VIH1 and VIH2, in Arabidopsis (Zhu et al., eLife, 2019). Jin obtained double knock-out Arabidopsis mutants for the diphosphoinositol pentakisphosphate kinases (PPIP5Ks) VIH1 and 2.
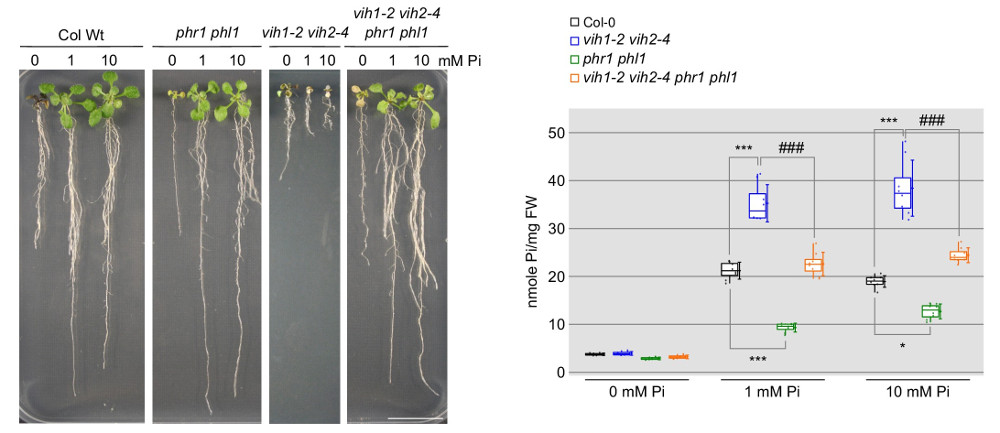
These mutant plants exhibit severe developmental defects and die at the seedling stage due to the constant expression of phosphate-starvation genes and excessive phosphate accumulation (Figure 11). Deleting phosphate starvation response transcription factors (PHR) partially rescues the vih1 vih2 mutant phenotypes (Figure 11), positioning these two kinases within the plant’s phosphate signaling pathway.
Jin and Kelvin went on to demonstrate that VIH1/2, like other PPIP5Ks, are bifunctional enzymes, consisting of a kinase domain that produces PP-InsPs and a phosphatase domain that breaks them down. The PP-InsP kinase domain generates 1,5-InsP8 from 5-InsP7 and has a Km for ATP that is close to the cytosolic concentration of ATP. Thus, 1-5-InsP8 synthesis is stimulated in Pi sufficient conditions, where ATP levels are high (Zhu et al., eLife, 2019). The phosphatase activity on the other hand is inhibited by inorganic phosphate (Zhu et al., eLife, 2019). This suggests that VIH1 and VIH2 can translate changes in cellular ATP and inorganic phosphate levels into adjustments in PP-InsP levels, enabling plants to monitor and maintain optimal phosphate balance.
Three different families of inositol pyrophosphate phosphatases contribute to PP-InsP catabolism and phosphate homeostasis.
Genetic rescue experiments with vih1 vih2 plants revealed that the PPIP5K kinase but not the phosphatase activity is essential in Arabidopsis (Zhu et al., eLife, 2019). This could be in part explained by the presence of other PP-InsP phosphatases that may act redundantly with the PPIP5K phosphatase domains of VIH1 and VIH2. PhD student Florian Laurent characterized NUDIX and PFA-DSP hydrolases as PP-InsP phosphatases in vitro (Laurent et al., PLoS Genet, 2024). Over-expression of NUDIX or PFA-DSPs in Arabidopsis indeed alters cellular PP-InsP pools, but potentially due to genetic redundancy no convincing loss-of-function phenotypes could be obtained (Laurent et al., PLoS Genet, 2024). Since fewer family members exist for both NUDIX and PFA-DSP hydrolases in Marchantia polymorpha,
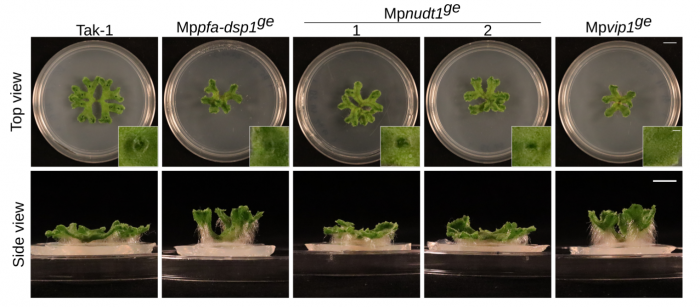

Florian and Felix Rico-Resendiz characterized CRISPR/Cas9 knock-out mutants of the different enzymes in Marchantia. Mppfa-dsp1 and Mpvip1 mutants indeed showed altered PP-InsP levels and associated growth phenotypes (Figure 12). A Mppfa-dsp1 Mpvip1 double mutant showed enhanced defects (Laurent et al., PLoS Genet, 2024). Phosphate levels and starvation responses are affected in the different mutants, suggesting that indeed several PP-InsP phosphatase enzyme families contribute to plant PP-InsP and Pi homeostasis.
Postdoctoral fellows Kitaik Lee and Pierre Raia next dissected the enzyme mechanism and regulation of the phosphatase domain in PPIP5Ks. They could show that the Vip1 phosphatase domain from baker’s yeast uses a conserved histidine phosphatase catalytic site to specifically hydrolyse 1,5-InsP8. The catalytic motif is structurally stabilized by a Zn-binding site. The enzyme uses a unique mechanism in which the PP-InsP substrate first binds to the outer surface of the enzyme and is then transported to the active site by a small GAF signaling domain (Figure 13). PPIP5K phosphatases are inhibited by inorganic phosphate (and sulfate) and appear to act independent from the N-terminal kinase module in the context of the full-length enzyme (Raia et al., Nat Commun, 2025).
SPX proteins target PHR transcription factors to control plant phosphate homeostasis
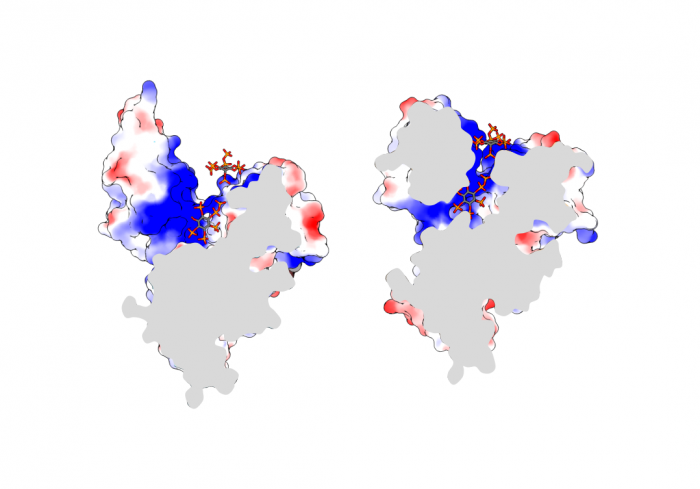
Previous studies had shown that stand-alone SPX proteins in Arabidopsis interact with PHR transcription factors, which activate the expression of phosphate starvation genes when phosphate is scarce (Puga et al., PNAS, 2014). The newly identified PP-InsP sensor function led us to investigate SPX-PHR interactions, which were found to be strictly dependent on PP-InsPs, particularly InsP8 (Wild et al., Science, 2016, Ried et al., Nat Commun, 2019). PhD student Rebekka Wild determined crystal structures of the Arabidopsis PHR1 coiled-coil domain and identified residues essential for interaction with the PP-InsP-bound SPX domain (Figure 14).
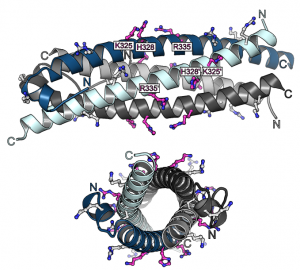
Postdoctoral fellow Martina Ried mutated these conserved interface residues and found that the mutations disrupted the interaction with the SPX receptor in vitro and in planta, resulting in constitutive phosphate starvation responses (Ried et al., Nat Commun, 2021).
In conjunction with our work on VIH1/2, we conclude that InsP8 regulates plant phosphate homeostasis by controlling the oligomeric state of PHRs and, consequently, their ability to bind promoters via their SPX receptors. We reconcile the earlier observations that SPX-PHR complexes are phosphate-dependent in plants by suggesting that, under low phosphate conditions, VIH1/2 kinases are likely deactivated, leading to a reduction in PP-InsP levels. This decline allows PHR transcription factors to bind to target promoters and activate phosphate response genes (Lorenzo-Orts et al., New Phytologist, 2020). The crystal structure of a rice SPX – PHR complex further supports this model (Guan et al., Nature Commun, 2022) (Figure 15).